The Human genome complete set of genetic information present in our cells consists of approximately 3 billion base pairs split among 23 pairs of chromosomes present in every cell of the body that encodes about 19000-20000 different kinds of proteins coding genes in a haploid cell. The human genome project has been completed in 2003 and the entire human genome has been sequenced. During the process of central dogma (DNA-RNA-Proteins) possible chances of errors in the process of encoding genes during transcription and translation processes.
Some repair mechanisms are available in cells for reducing errors but sometimes mutations occurred during DNA replication or RNA processing. Some mutations are less harmful and some may lead to serious problems in the functioning of the cell and prove dangerous for the body like cancer. Approximately, 90% of human genome variation comes due to change in single nucleotides position known as single nucleotide polymorphism (“snips”) such as in β-globin gene which ultimately leads to sickle cell anemia.
What is Cancer?
Cancer is an uncontrolled division of the cells and the most complex heterogeneous and dynamic disease. It is characterized as abnormal clones of cells that are caused by alteration of a genome or in gene duplication. Cancer can cause genetic and epigenetic alternation in the whole genome of the cell and activation of oncogenes. Epigenetic alteration changes in gene expression without alteration in the DNA sequence.
Hanahan and Weinberg in 2000 studied the functional logic of cancer. They described six possibilities during tumor development-sustaining proliferative signaling, evading growth suppressors, resisting cell death, enabling replicative immortality, inducing angiogenesis and activating invasion and metastasis.
One in eight deaths that occur due to cancer in the human population represents a major concern for our society, and the most efficient, as well as safe treatment of cancer, has become a major goal for researchers around the world. Genetic engineering techniques, especially recent advance therapy by CRISPR associated with Cas9 has become a pivotal role in the treatment of cancer.
Introduction to CRISPR Cas9
In 1987, Ishino et al first time discovered some unusual repeats of nucleotides in the genome of Escherichia coli with unknown functions. Later Mojica et al some of these types of repeats in other microbes termed as “Regularly Interspaced Short Palindromic Repeats or CRISPR”. It is a short nucleotides sequence present in the genome of Bacterial DNA which helps in the immune defense system for degrading foreign DNA or viral DNA by cutting with nucleases enzyme.
CRISPR associated Cas9 is an important endonuclease enzyme present in some bacteria (E. coli) which require guide RNA (gRNA) sequence for activation of both enzyme and selective target complementary DNA. CRISPR/Cas9 is less expensive, led to rapid expansion in the biomedical field, development of biotechnology products, treatment of genetic disease, cancer, targeting BCR-ABL and more specifically revolutionized technique used for genome editing. This article will provide an overview of current research studies and some of the future directions for the application of CRISPR/Cas9 technology in cancer treatment.
CRISPR/Cas9 System
CRISPR-short for clustered, Regularly Interspaced, Short Palindromic Repeats, is a genetic phenomenon found in microbes for chopping of DNA. When combined with certain proteins, typically one called Cas9 (endonuclease a scalpel-like enzyme), a biological complex that can cut and paste DNA as altering genetic code (Cas gene). And dead endonuclease Cas9 (dCas9) is the mutant form of Cas9 protein whose nuclease activity becomes diminished due to point mutation. The universal property of this system is that it required RNA molecules as guides. There are four major nuclease editing systems for producing double-standard break (DSB) in the cell from target DNA: –
- Zinc finger nucleases (ZFNs)
- Transcription activator-like effector nuclease (TALENs)
- Mega nucleases
- CRISPR/Cas9 nucleases
The only application of CRISPR/Cas9 involved guide RNA (gRNA) sequence which is complementary to target DNA. ZFNs, TALENs, and Mega nucleases required more time and costly procedure than CRISPR/Cas9. The gRNA binds with cas9 protein and then both are injected into target cells through vectors by using recombinant DNA technologies.
In 2013, Jennifer Doudna professor of California University, Berkeley and her colleagues developed the CRISPR-Cas9 gene expression system by fusing two RNA molecules into single guided RNA (gRNA). Case9 endonuclease consists of CRISPR RNA (crRNA) and transactivating CRISPR RNA (tracrRNA) to form gRNA. By manipulating the sequence of gRNA, the artificial Cas9 system can be prepared according to the target.
The desired sequence must have Protospacer adjacent motif (PAM) which is 2-6 base pair DNA sequence which is targeted by Cas9 nuclease. It is the component of foreign or viral DNA, not the bacterial CRISPR locus. After the generation of a double-standard break (DSB) at the target sequence, it can be repaired by non-homologous end-joining (NHEJ) resulting in little insertion or deletion (indels) in associated loss of functional gene (knockdown/knockout). And in the presence of donor DNA sequence, homology-directed recombination (HDR) mechanism performed by modifications at the target site which results in the gain of function (knock-in). NHEJ is mostly dominating during G1, S and G2 phase while HDR in late S and G2 phase. (Fig 1)
The working mechanism of CRISPR-Cas9
CRISPR/Cas9 is a defense system and immune machinery that bacteria used to attack viral invading DNA via CRISPR loci detection and further lead to the destruction of foreign genetic material. While the first-ever version of CRISPR technology engineered to work in human cells originated from bacteria Streptococcus Pyogenes.
As describes Cas9 nuclease consist of single guide RNA (sgRNA) contain both crRNA and tracrRNA. The activation of Cas9 nuclease depends upon the reorganization of the protospacer adjacent motif (PAM) which is present at the end of the target DNA sequence by sgRNA. PAM is a special marker for unwinding DNA only for target not it’s own. So, lack of PAM leads to the failure of unwinding DNA. After an unwinding of double-strand DNA from its upstream region, the cleavage process starts with the hybridization of sgRNA-DNA sequences. Then double standard breaks (DSBs) are produced by CRISPR-Cas9, triggered to NHEJ and HDR, leading to indels in target DNA. An enormous study shows that efficient knockdown or knockin in somatic cells by the CRISPR-Cas9 system.
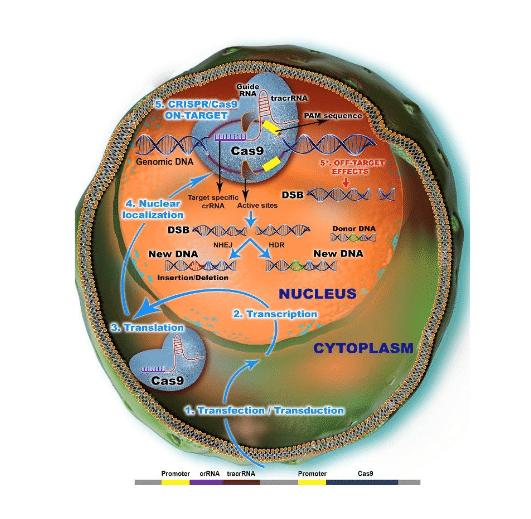
Figure 1. CRISPR/Cas9 Mechanism of Action
There are some delivery systems of CRISPR/Cas9 cancer gene therapy as adenoviruses (AVs) and adenosine associated viruses (AAVs) most widely used due to their high transfection efficiency. For the permanent expression, Cas9 and gRNA must be integrated into the host genome, the best option is to use Retrovirus. They have the capability of reverse transcriptase (RNA to DNA). Some others are fusion proteins of Cas9, Nano-formulations, and Membrane-derived vesicles.
CRISPR/Cas9 as Gene-editing tool for Cancer
After the discovery of the CRISPR/Cas9 genome editing tool in 2012, a publication demonstrated gene editing in Streptococcus Pyogenes. In this case, Cas9 protein which is responsible for genome editing binds with gRNA to specify the targeted DNA and cleavage result in the form of HDR and NHEJ strand natural pathways. Some gRNA of Cas9 only bind but do not start the process of cleavage. Usually used to stop the process of transcription (activator or repressor). CRISPR/Cas9 became the efficient tool of endogenous gene editing in mammalian cells.
First time in Streptococcus Pyogenes, a point mutation was expended in Cas9 protein for the formation of dCas9. dCas9 could be used for a variety of functions such as regulation of the transcriptional process, activator, and repressor with gRNA, editing of epigenetic marks and tag endogenous gene loci without change the host DNA sequence.
CRISPR first time applied to modulate somatic cancer-causing mutations in the liver of wild type mouse after also in human intestinal stem cells organoids by gene knockout of tumor suppressor genes PTEN and P53. Today this technique of gene repressor has been used for the elimination of lung, pancreatic and brain cell cancer. Sometimes, different kinds of point mutations have been induced as activation of KRAS and CTNNBI oncogenes.
CRISPR-based HDR can stimulate defined modifications in genetic sequences. A frameshift mutation in APC which is tumor repressor gene and oncogenic ALK-F1174L activating mutation has been repaired in colon cancer and neuroblastoma cells respectively. Some epigenetic drugs are also used extensively for the treatment of cancer but these have strong side-effects.
Now a day, many non-coding RNA (ncRNA) such as micro-RNA (miRNA) and short interfering RNA which are responsible for inhibition or activation of oncogenes, involves in progression and development of cancer could be suppressed by CRISPR/Cas9 technology (edit genetic switches). These technologies may have overcome the problems that can generate permanent modifications in somatic cells. In human and mouse-derived cells, authors found that huge chunks of DNA were unintentionally being deleted or rearranged so severely that cells may be lost its function about 15 percent cases.
The most important and widely clinical trials used by CRISPR cancer targeting and destruction are on Modified chimeric antigen receptor T-cells which is the most reliable technique than others. These modifications induced with Cas9-gRNA in T-cells known as CRISPR-based CAR T-cells and recently have been approved in the U.S and China for a clinical trial. This is based on knockout engineered programmed cell death protein-1 (PD-1) which is responsible for inhibition of T-cells receptor (Fig 2). The edited lymphocytes are added back into the patient. And four other trials have been done on different types of cancer which are based on gene knockout (PD-1) mechanism as prostate, bladder, esophageal and renal cell cancer.
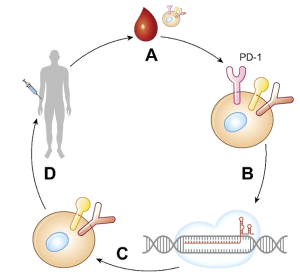
Figure 2 CRISPR/Cas9 as a therapeutic tool. Principle of first phase 1 clinical trials
Challenges and Future Aspects of CRISPR/Cas9
Today in cancer biology, CRISPR/Cas9 plays important role in the progression of cancer therapies and changed our perspective by providing new approaches for personalized therapy, immunotherapeutic applications, genetic disorder treatment, and etc. And this technology may accelerate gene therapies in future. Current work shows that CRISPR can correct gene mutations permanently in some animals. There are some hurdles of using in therapeutic applications of CRISPR method as mutation can be raised during NHEJ and HDR natural pathways. CRISPR/Cas9 has many off-target effects on DNA by generating DSBs at nonspecific sites in contrast with base editors in which chemically change one DNA base into another with an enzyme called deaminase.
To overcome this, a number of new techniques have been developed for avoiding off-target issues. The scientist has deployed a breakthrough CRISPR gene-editing tool to successfully multiple edits simultaneously to fragments of DNA extracted from a human cell. And recently researchers have developed a method for improving the accuracy of the CRISPR genome editing technology by an average of 50-fold by adds a short tail to the guide RNA which is used to identify a sequence of DNA for editing.
Conclusion
CRISPR-Cas9 is a promising tool for wiping out genetic mutations and precise methods of genetic editing that widely used in scientific research but it may be causing genetic damage of its own. CRISPR gene editing may be doing more damage than scientists though it may cut down some unwanted bits of genetic code. And researchers hope it could one day be used to selectively remove genes that result in medical problems such as HIV, sickle cell disease and cancer.
Read more about healthy living lifestyle
– If you are looking for guest posts in health “write for us” now.